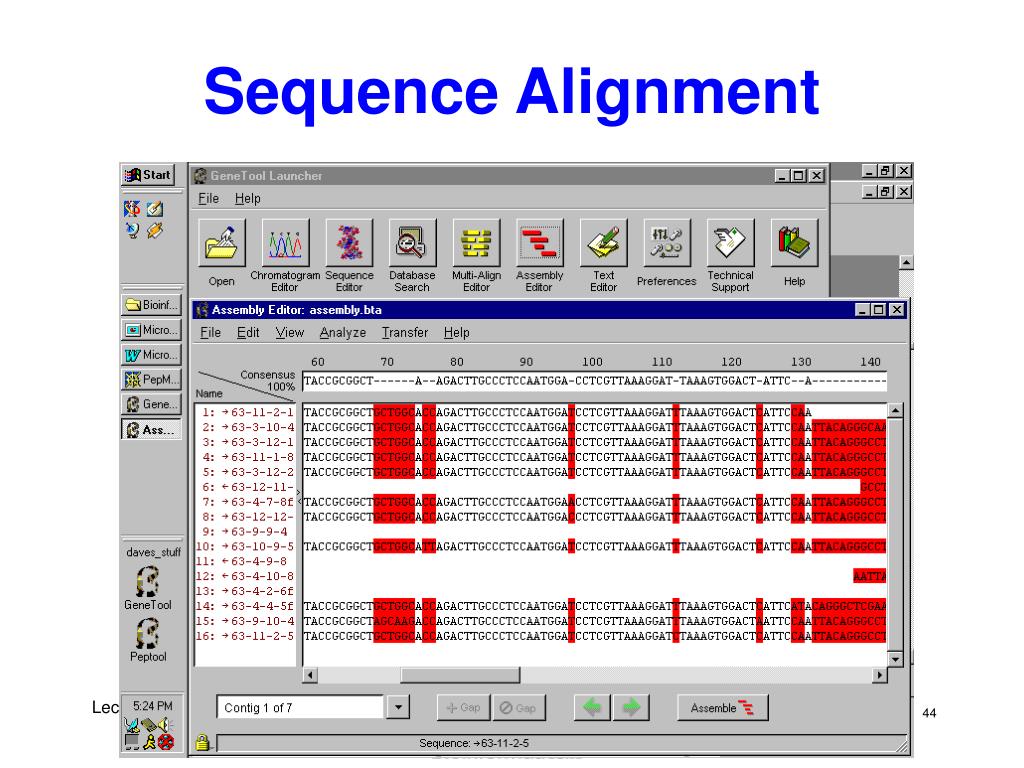
Here, we document different microbial sources of free DNA at the surface (0 to 200 m) versus depths of 250 to 1,000 m, suggesting that distinct free DNA production mechanisms may be present throughout the oligotrophic water column. The fate of this free DNA has both ecological consequences as a nutrient (N and P) source and potential evolutionary consequences as a source of genetic transformation.
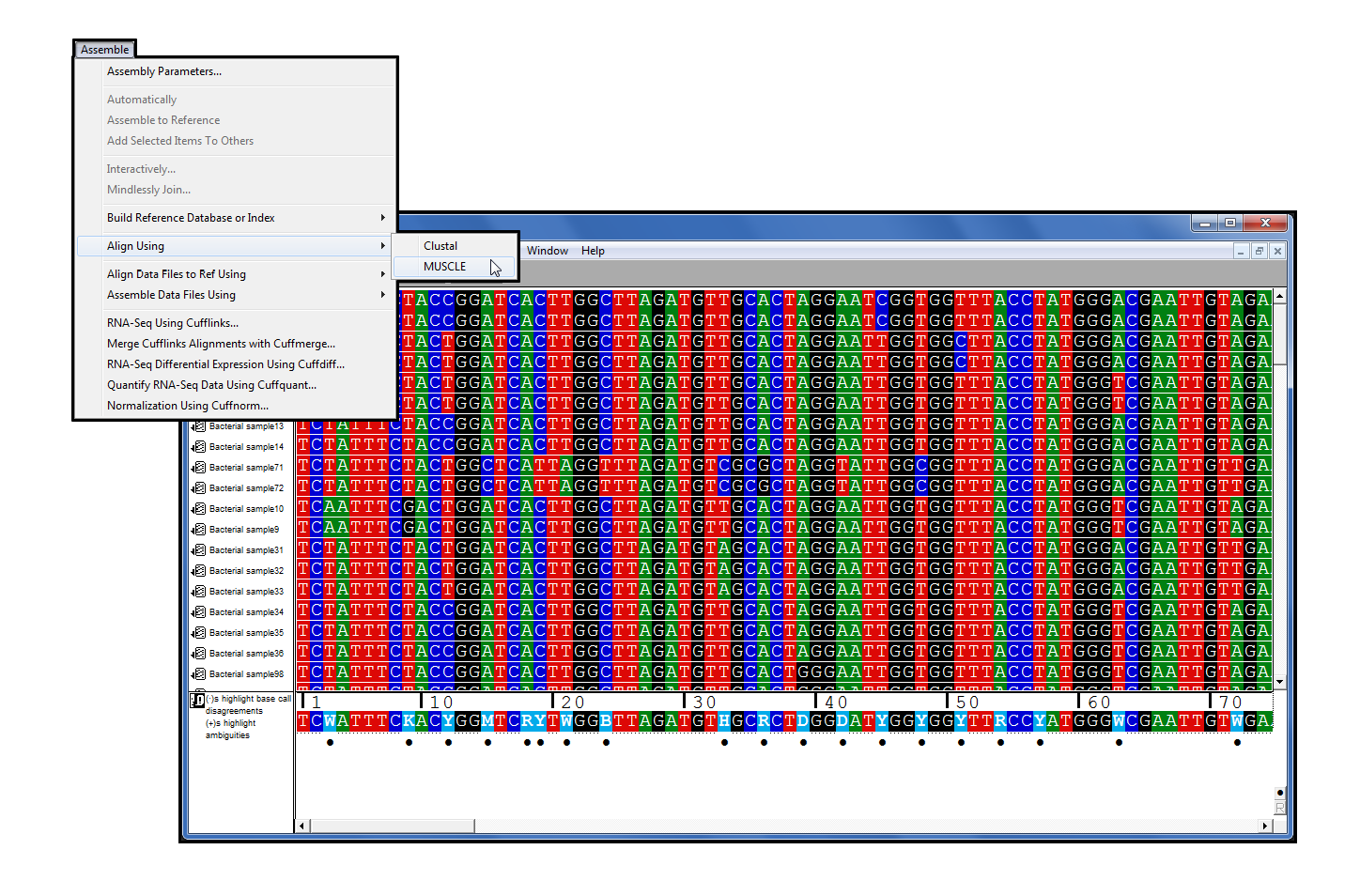
Here, we characterized exocellular free DNA via metagenomics, using a newly developed method that separates free DNA from cells, viruses, and vesicles, and facilitated the independent characterization of each fraction. IMPORTANCE With advances in metagenomic sequencing, the microbial composition of diverse environmental systems has been investigated, providing new perspectives on potential ecological dynamics and dimensions for experimental investigations. These results reveal the composition of free DNA in different regions of the water column (euphotic and mesopelagic zones), with implications for dissolved organic matter cycling and export (by way of sinking particles and/or migratory zooplankton) as a delivery mechanism. Throughout the water column, but especially in the mesopelagic zone, the composition of free DNA sequences was not always reflective of cooccurring microbial communities that inhabit the same sampling depth. A high proportion of mesopelagic zone free DNA sequences appeared to originate from surface waters, including a large amount of DNA contributed by high-light Prochlorococcus ecotypes. Euphotic zone free DNA (75 to 125 m) contained primarily bacterial and viral sequences, with bacteria dominating samples from the mesopelagic zone (500 to 1,000 m). Viral DNA consisted predominantly of myovirus-like and podovirus-like DNA and contained the highest proportion of unannotated sequences. Pelagibacter-like DNA dominated the vesicle fractions for all samples examined over a depth range of 75 to 500 m. Using a method that provides separation of these three fractions, we compared open ocean depth profiles of DNA associated with each fraction. It is composed of DNA-containing vesicles, viruses, and free DNA and is ubiquitous in all aquatic systems, although the sources, sinks, and ecological consequences are largely unknown. Edgar RC.Exocellular DNA is operationally defined as the fraction of the total DNA pool that passes through a membrane filter (0.1 μm). MUSCLE: a multiple sequence alignment method with reduced time and space complexity. MUSCLE: multiple sequence alignment with high accuracy and high throughput.
#Sequencher contig series
"Multiple sequence alignment with the Clustal series of programs". Chenna R, Sugawara H, Koike T, Lopez R, Gibson TJ, Higgins DG, Thompson JD (2003). Sequencher gives you access to MUSCLE’s power without the problems of learning to use the UNIX command line. MUSCLE is a command line program, which means that normally you would be using this program through a Terminal application. Use Sequencher’s tools to annotate your alignment or export your alignment and place it into a special phylogenetics program. Once the process is complete and MUSCLE has built the alignment, you will see the results as a contig within Sequencher. In the final two steps, the MUSCLE algorithm tries a number of different methods to see if it is possible to improve the tree and hence the multiple alignment. The second step uses this tree to guide a progressive alignment. The first step constructs a distance matrix between pairs of sequences using k-mer clustering, this is then converted into a tree. MUSCLE is said to have four major steps in its alignment process. Boasting both speed and accuracy, it compares very favorably to other multiple-sequence alignment programs. It joins Clustal, making it the second MSA program in Sequencher’s DNA-Seq Tools. MUSCLE, a multiple-sequence alignment (MSA) program, joins the Sequencher 5.1 family of plugins. Speed up your Clustal alignments by combining Clustal with the power of Sequencher's Assemble by Name functionality for aligning multiple sequences from different sources.įinally, export the results in a variety of different formats such as MSF, Phylip, NEXUS and FastA for use in other programs or simply create a consensus and export it. Once the alignment is complete you will see the results as a contig or contigs within Sequencher which you can subject to a variety of analyses. Choose from a range of parameters to control the alignment process. You can use Clustal to align your sequences directly from the Sequencher project window. It is a widely used multiple-sequence alignment program which works by determining all pairwise alignments on a set of sequences, then constructs a dendrogram grouping the sequences by approximate similarity and then finally performs the alignment using the dendogram as a guide. Clustal has been part of the Sequencher family of plugins since version 4.9.
